This week in MathOnco 199
Ecological theory, personalization, Genie genetic drift, predictive uncertainty, and more..
“This week in Mathematical Oncology” — Mar. 3, 2022
> mathematical-oncology.org
From the editor:
Today’s issue represents advances in topics such as ecological theory, personalization, genetic drift, predictive uncertainty, and more..
Jeffrey Westjeffrey.west@moffitt.org
Biallelic mutations in cancer genomes reveal local mutational determinants
Jonas Demeulemeester, Stefan C. Dentro, Moritz Gerstung, Peter Van LooMathematical model of hormone sensitive prostate cancer treatment using leuprolide: A small step towards personalization
Urszula Foryś, Alon Nahshony, Moran ElishmereniGenie: an interactive real-time simulation for teaching genetic drift
Andreina I. Castillo, Ben H. Roos, Michael S. Rosenberg, Reed A. Cartwright, Melissa A. WilsonEffective drug combinations in breast, colon and pancreatic cancer cells
Patricia Jaaks, Elizabeth A. Coker, Daniel J. Vis, Olivia Edwards, …, Andrea Bertotti, Livio Trusolino, Lodewyk Wessels & Mathew J. Garnett
Data-driven simulation of Fisher-Kolmogorov tumor growth models using Dynamic Mode Decomposition
Alex Viguerie, Malú Grave, Gabriel F. Barros, Guillermo Lorenzo, Alessandro Reali, Alvaro L.G.A. CoutinhoPredictive uncertainty in mechanistic models of cellular processes calibrated to experimental data
Michael Irvin, Arvind Ramanathan, Carlos F LopezTransmissible Cancer Evolution: The Under-Estimated Role of Environmental Factors in the “Perfect Storm” Theory
Sophie Tissot, Anne-Lise Gérard, Justine Boutry, Antoine M. Dujon, …, Rodrigo Hamede, Benjamin Roche, Beata Ujvari, Frédéric ThomasMathematical model of a cytokine storm
Irina Kareva, Faina Berezovskaya, Georgy KarevIndividual-based and continuum models of phenotypically heterogeneous growing cell populations
Fiona R Macfarlane, Xinran Ruan, Tommaso LorenziBorrowing ecological theory to infer interactions between sensitive and resistant breast cancer cell populations
Zachary Susswein, Surojeet Sengupta, Robert Clarke, Shweta Bansal
The Mathematical Oncology Blog — The “Behind the Paper” series
Rachel Cavill, Kateřina Staňková: “We decided we would like to understand how to get beyond models of cancer and its treatment that are fitted with volumetric information only, to be able to predict the effect of anti-cancer treatments and improve therapy through game-theoretic modeling.”Evolutionary therapy
Wikipedia
Evolutionary therapy has its own Wikipedia page now, thanks to Luka Opasic and Jake Scott. Check it out!
The newsletter now has a dedicated homepage where we post the cover artwork for each issue. We encourage submissions that coincide with the release of a recent paper from your group.
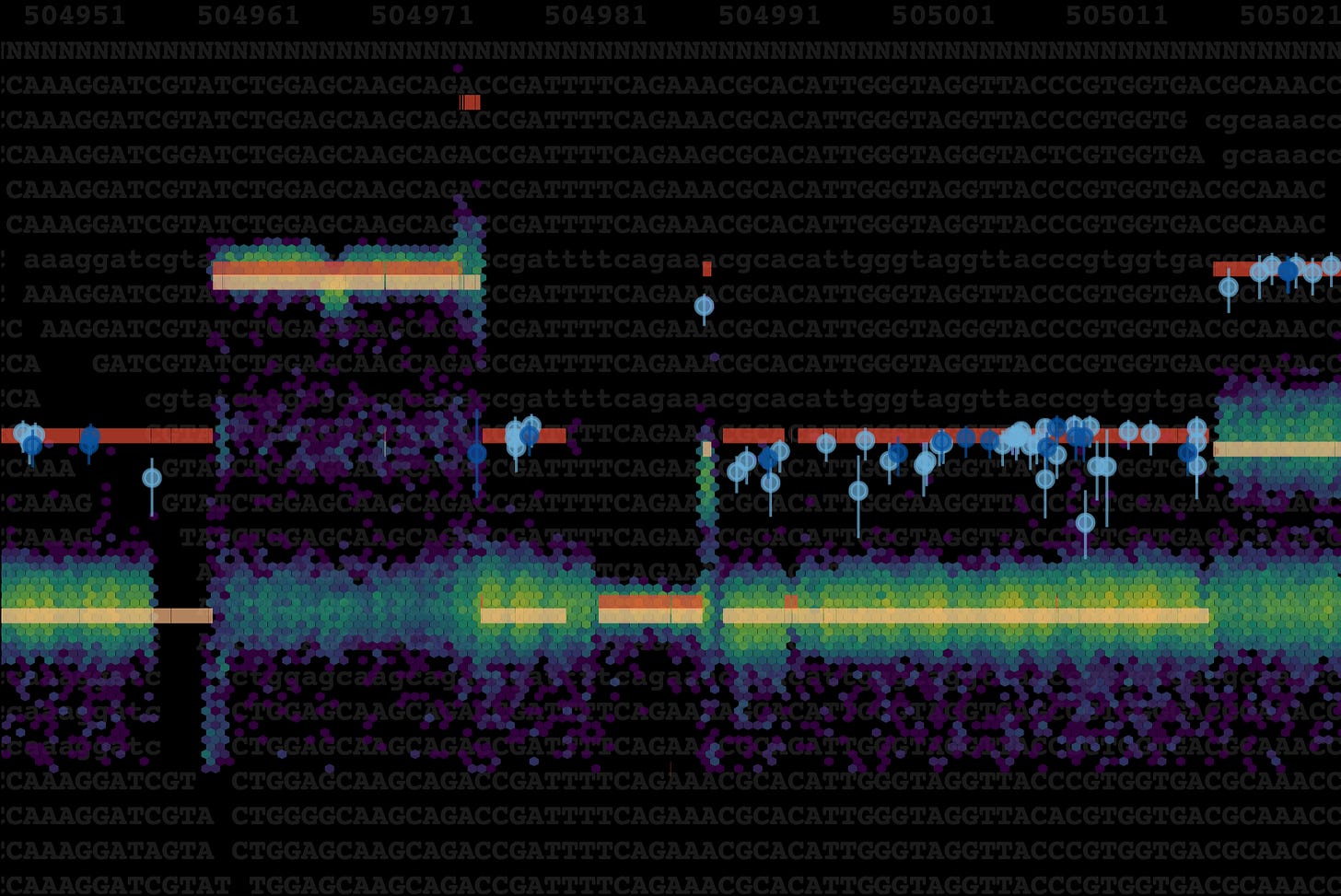
Caption: Do tumor cells sometimes make the same mistake twice? In our Nature Genetics paper, we identify biallelic mutations, where the same base is mutated independently on both parental alleles, across 2,658 cancer genomes and highlight the processes behind them. These mutations violate the infinite sites model of molecular evolution, a cornerstone of tumor phylogenetic analysis which is also often implied when calling, phasing and interpreting variants or studying the mutational landscape as a whole. The cover shows copy number estimates for chromosomes and mutations across part of a melanoma whole genome. Biallelic parallel mutations (blue circles) can be identified when their copy number is equal to the total copy number (red bars) in heterozygous regions of the tumor genome.
Created by: Jonas Demeulemeester (@zeunas & @VanLooLab)
Visit the mathematical oncology page to view jobs, meetings, and special issues. We will post new additions here, but the full list can found at mathematical-oncology.org.
1. Jobs
Graduate Summer Intern in Machine Learning, Pfizer Oncology La Jolla (or remote), CA (Blerta Shtylla, Email: Blerta.Shtylla@pfizer.com or Kamrine Poels Email: Kamrine.Poels@pfizer.com ), please contract directly if interested.
Current subscriber count: >1.1k